Last time we investigated the k-nearest-neighbors algorithm and the underlying idea that one can learn a classification rule by copying the known classification of nearby data points. This required that we view our data as sitting inside a metric space; that is, we imposed a kind of geometric structure on our data. One glaring problem is that there may be no reasonable way to do this. While we mentioned scaling issues and provided a number of possible metrics in our primer, a more common problem is that the data simply isn’t numeric.
For instance, a poll of US citizens might ask the respondent to select which of a number of issues he cares most about. There could be 50 choices, and there is no reasonable way to assign these numerical values so that all are equidistant in the resulting metric space.
Another issue is that the quality of the data could be bad. For instance, there may be missing values for some attributes (e.g., a respondent may neglect to answer one or more questions). Alternatively, the attributes or the classification label could be wrong; that is, the data could exhibit noise. For k-nearest-neighbors, missing an attribute means we can’t (or can’t accurately) compute the distance function. And noisy data can interfere with our choice of $ k$. In particular, certain regions might be better with a smaller value of $ k$, while regions with noisier data might require a larger $ k$ to achieve the same accuracy rate. Without making the algorithm sufficiently more complicated to vary $ k$ when necessary, our classification accuracy will suffer.
In this post we’ll see how decision trees can alleviate these issues, and we’ll test the decision tree on an imperfect data set of congressional voting records. We’ll implement the algorithm in Python, and test it on the problem of predicting the US political party affiliation of members of Congress based on their votes for a number of resolutions. As usual, we post the entire code and data set on this blog’s Github page.
Before going on, the reader is encouraged to read our primer on trees. We will assume familiarity with the terminology defined there.
Intuition
Imagine we have a data set where each record is a list of categorical weather conditions on a randomly selected number of days, and the labels correspond to whether a girl named Arya went for a horse ride on that day. Let’s also assume she would like to go for a ride every day, and the only thing that might prohibit her from doing so is adverse weather. In this case, the input variables will be the condition in the sky (sunny, cloudy, rainy, and snow), the temperature (cold, warm, and hot), the relative humidity (low, medium, and high), and the wind speed (low and high). The output variable will be whether Arya goes on a horse ride that day. Some entries in this data set might look like:
Arya's Riding Data
Sky Temperature Humidity Wind Horse Ride
Cloudy Warm Low Low Yes
Rainy Cold Medium Low No
Sunny Warm Medium Low Yes
Sunny Hot High High No
Snow Cold Low High No
Rainy Warm High Low Yes
In this case, one might reasonably guess that certain weather features are more important than others in determining whether Arya can go for a horse ride. For instance, the difference between sun and rain/snow should be a strong indicator, although it is not always correct in this data set. In other words, we’re looking for a weather feature that best separates the data into its respective classes. Of course, we’ll need a rigorous way to measure how good that separation is, but intuitively we can continue.
For example, we might split based on the wind speed feature. In this case, we have two smaller data sets corresponding to the entries where the wind is high and low. The corresponding table might look like:
Arya's Riding Data, Wind = High
Sky Temperature Humidity Horse Ride
Sunny Hot High No
Snow Cold Low No
Arya's Riding Data, Wind = Low
Sky Temperature Humidity Horse Ride
Cloudy Warm Low Yes
Rainy Cold Medium No
Sunny Warm Medium Yes
Rainy Warm High Yes
In this case, Arya is never known to ride a horse when the wind speed is high, and there is only one occasion when she doesn’t ride a horse and the wind speed is low. Taking this one step further, we can repeat the splitting process on the “Wind = Low” data in search of a complete split between the two output classes. We can see by visual inspection that the only “no” instance occurs when the temperature is cold. Hence, we should split on the temperature feature.
It is not useful to write out another set of tables (one feels the pain already when imagining a data set with a thousand entries), because in fact there is a better representation. The astute reader will have already recognized that our process of picking particular values for the weather features is just the process of traversing a tree.
Let’s investigate this idea closer. Imagine we have a tree where the root node corresponds to the Wind feature, and it has two edges connected to child nodes; one edge corresponds to the value “Low” and the other to “High.” That is, the process of traveling from the root to a child along an edge is the process of selecting only those data points whose “Wind” feature is that edge’s label. We can take the child corresponding to “Low” wind and have it represent the Temperature feature, further adding three child nodes with edges corresponding to the “Cold,” “Warm,” and “Hot” values.
We can stop this process once the choice of features completely splits our data set. Pictorially, our tree would look like this:
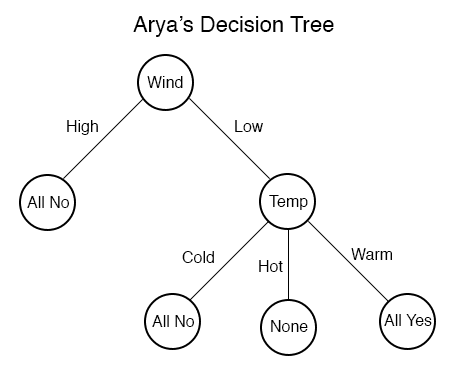
We reasonably decide to stop the traversal when all of the examples in the split are in the same class. More so, we would not want to include the option for the temperature to be Hot in the right subtree, because the data tells us nothing about such a scenario (as indicated by the “None” in the corresponding leaf).
Now that we have the data organized as a tree, we can try to classify new data with it. Suppose the new example is:
Sky Temperature Humidity Wind Horse Ride
Rainy Cold Low Low ?
We first inspect the wind speed feature, and seeing that it is “Low,” we follow the edge to the right subtree and repeat. Seeing that the temperature feature is “Cold,” we further descend down the “Cold” branch, reaching the “All No” leaf. Since this leaf corresponds to examples we’ve seen which are all in the “No” class, we should classify the new data as “No” as well.
Summarizing, given a new piece of data, we can traverse the tree according to the values of its features until we reach a leaf node. If the leaf node is “All No,” then we classify the new set of weather conditions as a “No,” and if it is “All Yes,” we classify as “Yes.”
Of course, this tree makes it clear that this toy data set is much too small to be useful, and the rules we’ve extrapolated from it are ridiculous. In particular, surely some people ride horses when the wind speed is high, and they would be unlikely to do so in a warm, low-wind thunderstorm. Nevertheless, we might expect a larger data set to yield a more sensible tree, as the data would more precisely reflect the true reasons one might refrain from riding a hose.
Before we generalize this example to any data set, we should note that there is yet another form for our classification rule. In particular, we can write the traversal from the root to the rightmost leaf as a boolean expression of the form:
$$\displaystyle \textup{Wind = “Low”} \wedge \textup{Temp = “Warm”}$$An example will be classified as “Yes” if and only if the wind feature is “High” and the temperature feature is “Warm” (here the wedge symbol $ \wedge$ means “and,” and is called a conjunction). If we had multiple such routes leading to leaves labeled with “All Yes,” say a branch for wind being “High” and sky being “Sunny,” we could expand our expression as a disjunction (an “or,” denoted $ \vee$) of the two corresponding conjunctions as follows:
$ \displaystyle (\textup{Wind = “Low”} \wedge \textup{Temp = “Warm”}) \vee (\textup{Wind = “High”} \wedge \textup{Sky = “Sunny”})$
In the parlance of formal logic, this kind of expression is called the disjunctive normal form, that is, a disjunction of conjunctions. It’s an easy exercise to prove that every propositional statement (in our case, using only and, or, and parentheses for grouping) can be put into disjunctive normal form. That is, any boolean condition that can be used to classify the data can be expressed in a disjunctive normal form, and hence as a decision tree.
Such a “boolean condition” is an example of a hypothesis, which is the formal term for the rule an algorithm uses to classify new data points. We call the set of all possible hypotheses expressible by our algorithm the hypothesis space of our algorithm. What we’ve just shown above is that decision trees have a large and well-defined hypothesis space. On the other hand, it is much more difficult to describe the hypothesis space for an algorithm like k-nearest-neighbors. This is one argument in favor of decision trees: they have a well-understood hypothesis space, and that makes them analytically tractable and interpretable.
Using Entropy to Find Optimal Splits
The real problem here is not in using a decision tree, but in constructing one from data alone. At any step in the process we outlined in the example above, we need to determine which feature is the right one to split the data on. That is, we need to choose the labels for the interior nodes in so that the resulting data subsets are as homogeneous as possible. In particular, it would be nice to have a quantitative way to measure the quality of a split. Then at each step we could simply choose the feature whose split yields the highest value under this measurement.
While we won’t derive such a measurement in this post, we will use one that has an extensive history of applications: Shannon entropy.
Definition: Let $ D$ be a discrete probability distribution $ (p_1, p_2, \dots, p_n)$. Then the Shannon entropy of $ D$, denoted $ E(p_1, \dots, p_n)$ is
$$\displaystyle E(p_1, \dots , p_n) = – \sum_{i=1}^n p_i \log(p_i)$$Where the logarithms are taken in base 2.
In English, there are $ n$ possible outcomes numbered 1 to $ n$, and the probability that an instance drawn from $ D$ results in the outcome $ k$ is $ p_k$. Then Shannon’s entropy function computes a numerical quantity describing how “dispersed” the outcomes are.
While there are many other useful interpretations of Shannon entropy, we only need it to describe how well the data is split into its classes. For our purposes, the probability distribution will simply be the observed proportions of data with respect to their class labels. In the case of Arya’s horse riding, the initial distribution would be $ (1/2, 1/2)$, giving an entropy of $ 1$.
Let’s verify that Shannon’s entropy function makes sense for our problem. Specifically, the best scenario for splitting the data on a feature is a perfect split; that is, each subset only has data from one class. On the other hand, the worst case would be where each subset is uniformly distributed across all classes (if there are $ n$ classes, then each subset has $ 1/n$ of its data from each class).
Indeed, if we adopt the convention that $ \log(0) = 0$, then the entropy of $ (1,0, \dots, 0)$ consists of a single term $ -1 \log(1) = 0$. It is clear that this does not depend on the position of the 1 within the probability distribution. On the other hand, the entropy of $ (1/n, \dots, 1/n)$ is
$$\displaystyle -\sum_{i=1}^n\frac{1}{n}\log \left (\frac{1}{n} \right ) = -\log \left (\frac{1}{n} \right ) = -(0 – \log(n)) = \log(n)$$A well-known property of the entropy function tells us that this is in fact the maximum value for this function.
Summarizing this, in the best case entropy is minimized after the split, and in the worst case entropy is maximized. But we can’t simply look at the entropy of each subset after splitting. We need a sensible way to combine these entropies and to compare them with the entropy of the data before splitting. In particular, we would quantify the “decrease” in entropy caused by a split, and maximize that quantity.
Definition: Let $ S$ be a data set and $ A$ a feature with values $ v \in V$, and let $ E$ denote Shannon’s entropy function. Moreover, let $ S_v$ denote the subset of $ S$ for which the feature $ A$ has the value $ v$. The gain of a split along the feature $ A$, denoted $ G(S,A)$ is
$$\displaystyle G(S,A) = E(S) – \sum_{v \in V} \frac{|S_v|}{|S|} E(S_v)$$That is, we are taking the difference of the entropy before the split, and subtracting off the entropies of each part after splitting, with an appropriate weight depending on the size of each piece. Indeed, if the entropy grows after the split (that is if the data becomes more mixed), then this number will be small. On the other hand if the split separates the classes nicely, each subset $ S_v$ will have small entropy, and hence the value will be large.
It requires a bit of mathematical tinkering to be completely comfortable that this function actually does what we want it to (for instance, it is not obvious that this function is non-negative; does it make sense to have a negative gain?). We won’t tarry in those details (this author has spent at least a day or two ironing them out), but we can rest assured that this function has been studied extensively, and nothing unexpected happens.
So now the algorithm for building trees is apparent: at each stage, simply pick the feature for which the gain function is maximized, and split the data on that feature. Create a child node for each of the subsets in the split, and connect them via edges with labels corresponding to the chosen feature value for that piece.
This algorithm is classically called ID3, and we’ll implement it in the next section.
Building a Decision Tree in Python
As with our primer on trees, we can use a quite simple data structure to represent the tree, but here we need a few extra pieces of data associated with each node.
class Tree:
def __init__(self, parent=None):
self.parent = parent
self.children = []
self.splitFeature = None
self.splitFeatureValue = None
self.label = None
In particular, now that features can have more than two possible values, we need to allow for an arbitrarily long list of child nodes. In addition, we add three pieces of data (with default values None): the splitFeature is the feature for which each of its children assumes a separate value; the splitFeatureValue is the feature assumed for its parent’s split; and the label (which is None for all interior nodes) is the final classification label for a leaf.
We also need to nail down our representations for the data. In particular, we will represent a data set as a list of pairs of the form (point, label), where the point is itself a list of the feature values, and the label is a string.
Now given a data set the first thing we need to do is compute its entropy. For that we can first convert it to a distribution (in the sense defined above, a list of probabilities which sum to 1):
def dataToDistribution(data):
''' Turn a dataset which has n possible classification labels into a
probability distribution with n entries. '''
allLabels = [label for (point, label) in data]
numEntries = len(allLabels)
possibleLabels = set(allLabels)
dist = []
for aLabel in possibleLabels:
dist.append(float(allLabels.count(aLabel)) / numEntries)
return dist
And we can compute the entropy of such a distribution in the obvious way:
def entropy(dist):
''' Compute the Shannon entropy of the given probability distribution. '''
return -sum([p * math.log(p, 2) for p in dist])
Now in order to compute the gain of a data set by splitting on a particular value, we need to be able to split the data set. To do this, we identify features with their index in the list of feature values of a given data point, enumerate all possible values of that feature, and generate the needed subsets one at a time. In particular, we use a Python generator object:
def splitData(data, featureIndex):
''' Iterate over the subsets of data corresponding to each value
of the feature at the index featureIndex. '''
# get possible values of the given feature
attrValues = [point[featureIndex] for (point, label) in data]
for aValue in set(attrValues):
dataSubset = [(point, label) for (point, label) in data
if point[featureIndex] == aValue]
yield dataSubset
So to compute the gain, we simply need to iterate over the set of all splits, and compute the entropy of each split. In code:
def gain(data, featureIndex):
''' Compute the expected gain from splitting the data along all possible
values of feature. '''
entropyGain = entropy(dataToDistribution(data))
for dataSubset in splitData(data, featureIndex):
entropyGain -= entropy(dataToDistribution(dataSubset))
return entropyGain
Of course, the best split (represented as the best feature to split on) is given by such a line of code as:
bestFeature = max(range(n), key=lambda index: gain(data, index))
We can’t quite use this line exactly though, because while we’re building up the decision tree (which will of course be a recursive process) we need to keep track of which features have been split on previously and which have not; this data is different for each possible traversal of the tree. In the end, our function to build a decision tree requires three pieces of data: the current subset of the data to investigate, the root of the current subtree that we are in the process of building, and the set of features we have yet to split on.
Of course, this raises the obvious question about the base cases. One base case will be when we run out of data to split; that is, when our input data all have the same classification label. To check for this we implement a function called “homogeneous”
def homogeneous(data):
''' Return True if the data have the same label, and False otherwise. '''
return len(set([label for (point, label) in data])) <= 1
The other base case is when we run out of good features to split on. Of course, if the true classification function is actually a decision tree then this won’t be the case. But now that we’re in the real world, we can imagine there may be two data points with identical features but different classes. Perhaps the simplest way to remedy this situation is to terminate the tree at that point (when we run out of features to split on, or no split gives positive gain), and use a simple majority vote to label the new leaf. In a sense, this strategy is a sort of nearest-neighbors vote as a default. To implement this, we have a function which simply patches up the leaf appropriately:
def majorityVote(data, node):
''' Label node with the majority of the class labels in the given data set. '''
labels = [label for (pt, label) in data]
choice = max(set(labels), key=labels.count)
node.label = choice
return node
The base cases show up rather plainly in the code to follow, so let us instead focus on the inductive step. We declare our function to accept the data set in question, the root of the subtree to be built, and a list of the remaining allowable features to split on. The function begins with:
def buildDecisionTree(data, root, remainingFeatures):
''' Build a decision tree from the given data, appending the children
to the given root node (which may be the root of a subtree). '''
if homogeneous(data):
root.label = data[0][1]
return root
if len(remainingFeatures) == 0:
return majorityVote(data, root)
# find the index of the best feature to split on
bestFeature = max(remainingFeatures, key=lambda index: gain(data, index))
if gain(data, bestFeature) == 0:
return majorityVote(data, root)
root.splitFeature = bestFeature
Here we see the base cases, and the selection of the best feature to split on. As a side remark, we observe this is not the most efficient implementation. We admittedly call the gain function and splitData functions more often than necessary, but we feel what is lost in runtime speed is gained in code legibility.
Once we bypass the three base cases, and we have determined the right split, we just do it:
def buildDecisionTree(data, root, remainingFeatures):
''' Build a decision tree from the given data, appending the children
to the given root node (which may be the root of a subtree). '''
...
# add child nodes and process recursively
for dataSubset in splitData(data, bestFeature):
aChild = Tree(parent=root)
aChild.splitFeatureValue = dataSubset[0][0][bestFeature]
root.children.append(aChild)
buildDecisionTree(dataSubset, aChild, remainingFeatures - set([bestFeature]))
return root
Here we iterate over the subsets of data after the split, and create a child node for each. We then assign the child its corresponding feature value in the splitFeatureValue variable, and append the child to the root’s list of children. Next is where we first see the remainingFeatures set come into play, and in particular we note the overloaded minus sign as an operation on sets. This is a feature of python sets, and in particular it behaves exactly like set exclusion in mathematics. The astute programmer will note that the minus operation generates a *new set, so that further recursive calls to buildDecisionTree will not be affected by what happens in this recursive call.
Now the first call to this function requires some initial parameter setup, so we define a convenience function that only requires a single argument: the data.
def decisionTree(data):
return buildDecisionTree(data, Tree(), set(range(len(data[0][0]))))
Classifying New Data
The last piece of the puzzle is to classify a new piece of data once we’ve constructed the decision tree. This is a considerably simpler recursive process. If the current node is a leaf, output its label. Otherwise, recursively search the subtree (the child of the current node) whose splitFeatureValue matches the new data’s choice of the feature being split. In code,
def classify(tree, point):
''' Classify a data point by traversing the given decision tree. '''
if tree.children == []:
return tree.label
else:
matchingChildren = [child for child in tree.children
if child.splitFeatureValue == point[tree.splitFeature]]
return classify(matchingChildren[0], point)
And we can use this function to naturally test a dataset:
def testClassification(data, tree):
actualLabels = [label for point, label in data]
predictedLabels = [classify(tree, point) for point, label in data]
correctLabels = [(1 if a == b else 0) for a,b in zip(actualLabels, predictedLabels)]
return float(sum(correctLabels)) / len(actualLabels)
But now we run into the issue of noisy data. What if one wants to classify a point where one of the feature values which is used in the tree is unknown? One can take many approaches to remedy this, and we choose a simple one: simply search both routes, and use a majority vote when reaching a leaf. This requires us to add one additional piece of information to the leaf nodes: the total number of labels in each class used to build that leaf (recall, one of our stopping conditions resulted in a leaf having heterogeneous data). We omit the details here, but the reader is invited to read them on this blog’s Github page, where as usual we have provided all of the code used in this post.
Classifying Political Parties Based on Congressional Votes
We now move to a concrete application of decision trees. The data set we will work with comes from the UCI machine learning repository, and it records the votes cast by the US House of Representatives during a particular session of Congress in 1984. The data set has 16 features; that is, there were 16 different measures considered “key” measures that were vote upon during this session. So each point in the dataset represents the 16 votes of a single House member in that session. With a bit of reformatting, we provide the complete data set on this blog’s Github page.
Our goal is to learn party membership based on the voting records. This data set is rife with missing values; roughly half of the members abstained from voting on some of these measures. So we constructed a decision tree from the clean portion of the data, and use that to classify the remainder of the data.
Indeed, this data fits precisely into the algorithm we designed above. The code to construct a tree is almost trivial:
with open('house-votes-1984.txt', 'r') as inputFile:
lines = inputFile.readlines()
data = [line.strip().split(',') for line in lines]
data = [(x[1:], x[0]) for x in data]
cleanData = [x for x in data if '?' not in x[0]]
noisyData = [x for x in data if '?' in x[0]]
tree = decisionTree(cleanData)
Indeed, the classification accuracy in doing this is around 90%. In addition (though we will revisit the concept of overfitting later), this is stable despite variation in the size of the subset of data used to build the tree. The graph below shows this.
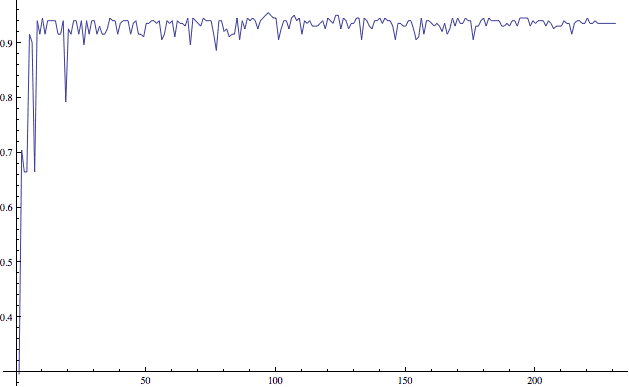
Inspecting the trees generated in this process, it appears that the most prominent feature to split on is the adoption of a new budget resolution. Very few Demorats voted in favor of this, so for many of the random subsets of the data, a split on this feature left one side homogeneously Republican.
Overfitting, Pruning, and Other Issues
Now there are some obvious shortcomings to the method in general. If the data set used to build the decision tree is enormous (in dimension or in number of points), then the resulting decision tree can be arbitrarily large in size. In addition, there is the pitfall of overfitting to this particular data set. For the party classification problem above, the point is to extend the classification to any population of people who might vote on these issues (or, more narrowly, to any politician who might vote on these issues). If we make our decision tree very large, then the hypothesis may be overly specific to the people in the sample used, and hence will not generalize well.
This problem is called overfitting to the data, and it’s a prevalent concern among all machine learning algorithms. There are a number of ways to avoid it for decision trees. Perhaps the most common is the idea of pruning: one temporarily removes all possible proper subtrees and reevaluates the classification accuracy for that removal. Whichever subtree results in the greatest increase in accuracy is actually removed, and it is replaced with a single leaf whose label corresponds to the majority label of the data points used to create the entire subtree. This process is then repeated until there are no possible improvements, or the gain is sufficiently small.
From a statistical point of view one could say this process is that of ignoring outliers. Any points which do not generally agree with the whole trend of the data set (hence, create their own branches in the decision tree) are simply removed. From a theoretical point of a view, a smaller decision tree satisfies the principle of Occam’s razor: a simpler hypothesis is more accurate by virtue of being simple.
While we won’t implement a pruning method here (indeed, we didn’t detect any overfitting in the congressional voting example), but it would be a nice exercise for the reader to wet his feet with the code given above. Finally, there are other algorithms to build decision trees that we haven’t mentioned here. You can see a list of such algorithms on the relevant wikipedia page. Because of the success of ID3, there is a large body of research on such algorithms.
In any event, next time we’ll investigate yet another machine learning method: that of neural networks. We’ll also start to look at more general frameworks for computational learning theory. That is, we’ll exercise the full might of theoretical mathematics to reason about how hard certain problems are to learn (or whether they can be learned at all).
Until then!
Want to respond? Send me an email, post a webmention, or find me elsewhere on the internet.